

Available from Reaxense
This protein is integrated into the Receptor.AI ecosystem as a prospective target with high therapeutic potential. We performed a comprehensive characterization of Histone H3.3 including:
1. LLM-powered literature research
Our custom-tailored LLM extracted and formalized all relevant information about the protein from a large set of structured and unstructured data sources and stored it in the form of a Knowledge Graph. This comprehensive analysis allowed us to gain insight into Histone H3.3 therapeutic significance, existing small molecule ligands, relevant off-targets, and protein-protein interactions.
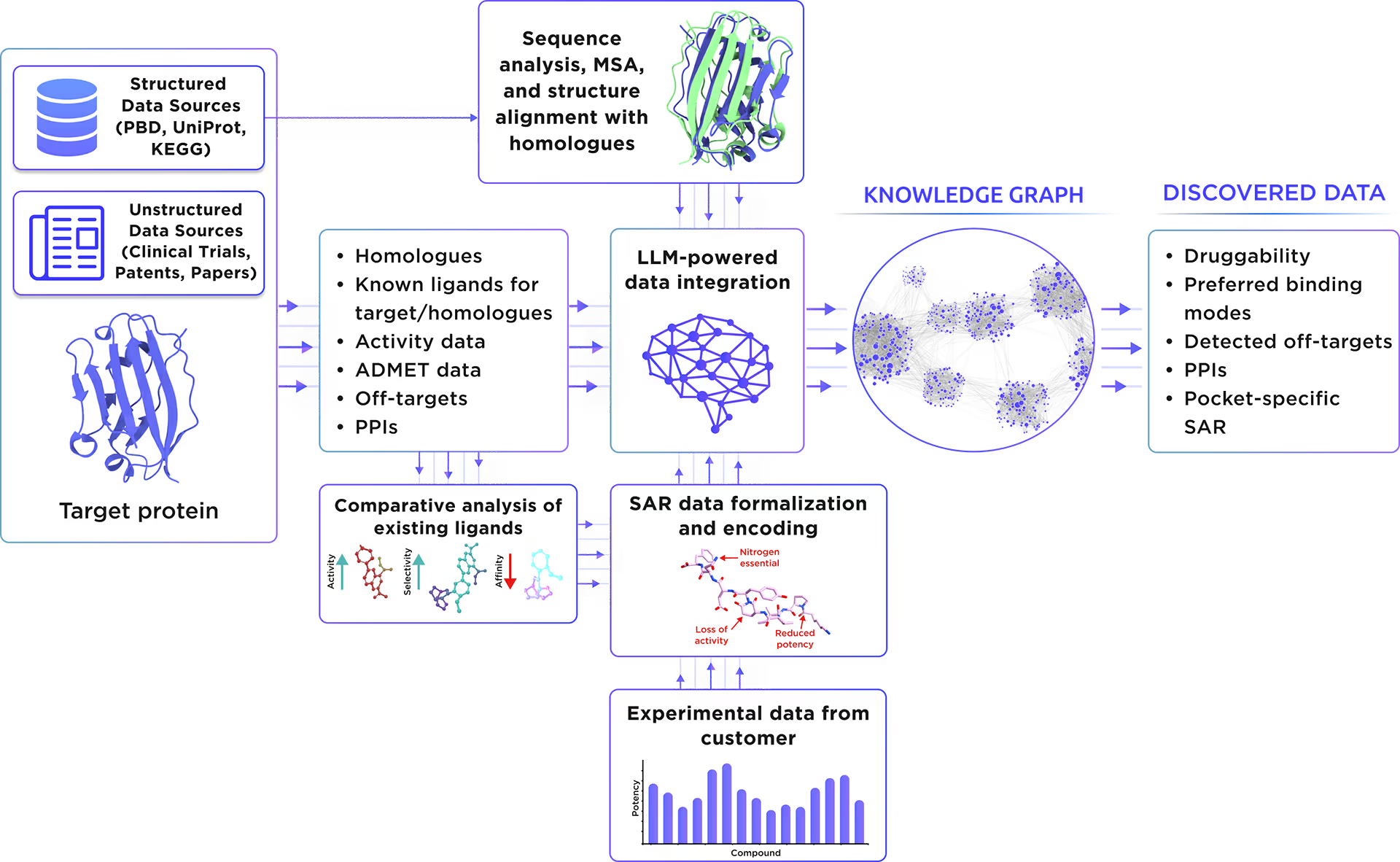
Fig. 1. Preliminary target research workflow
2. AI-Driven Conformational Ensemble Generation
Starting from the initial protein structure, we employed advanced AI algorithms to predict alternative functional states of Histone H3.3, including large-scale conformational changes along "soft" collective coordinates. Through molecular simulations with AI-enhanced sampling and trajectory clustering, we explored the broad conformational space of the protein and identified its representative structures. Utilizing diffusion-based AI models and active learning AutoML, we generated a statistically robust ensemble of equilibrium protein conformations that capture the receptor's full dynamic behavior, providing a robust foundation for accurate structure-based drug design.
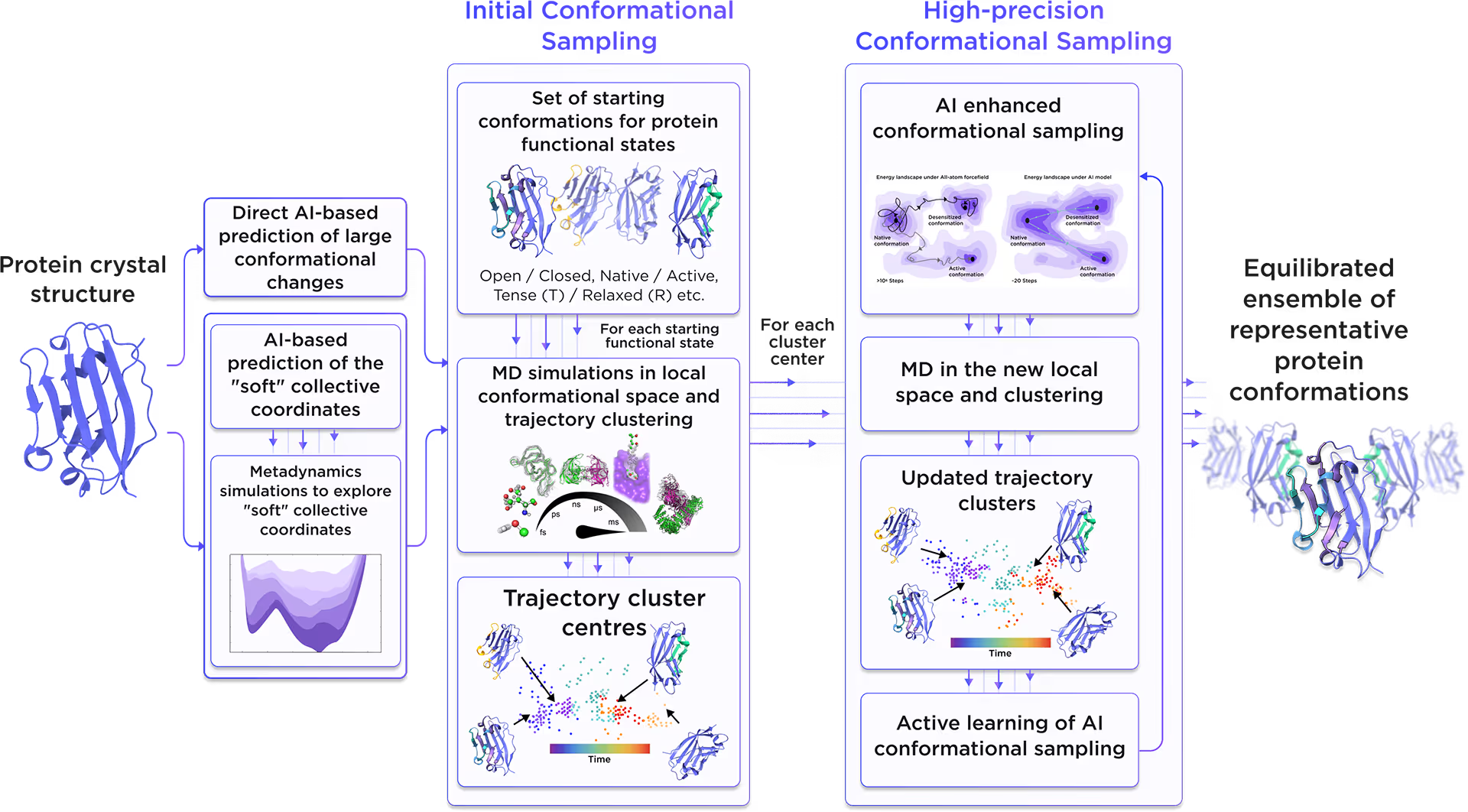
Fig. 2. AI-powered molecular dynamics simulations workflow
3. Binding pockets identification and characterization
We employed the AI-based pocket prediction module to discover orthosteric, allosteric, hidden, and cryptic binding pockets on the protein’s surface. Our technique integrates the LLM-driven literature search and structure-aware ensemble-based pocket detection algorithm that utilizes previously established protein dynamics. Tentative pockets are then subject to AI scoring and ranking with simultaneous detection of false positives. In the final step, the AI model assesses the druggability of each pocket enabling a comprehensive selection of the most promising pockets for further targeting.
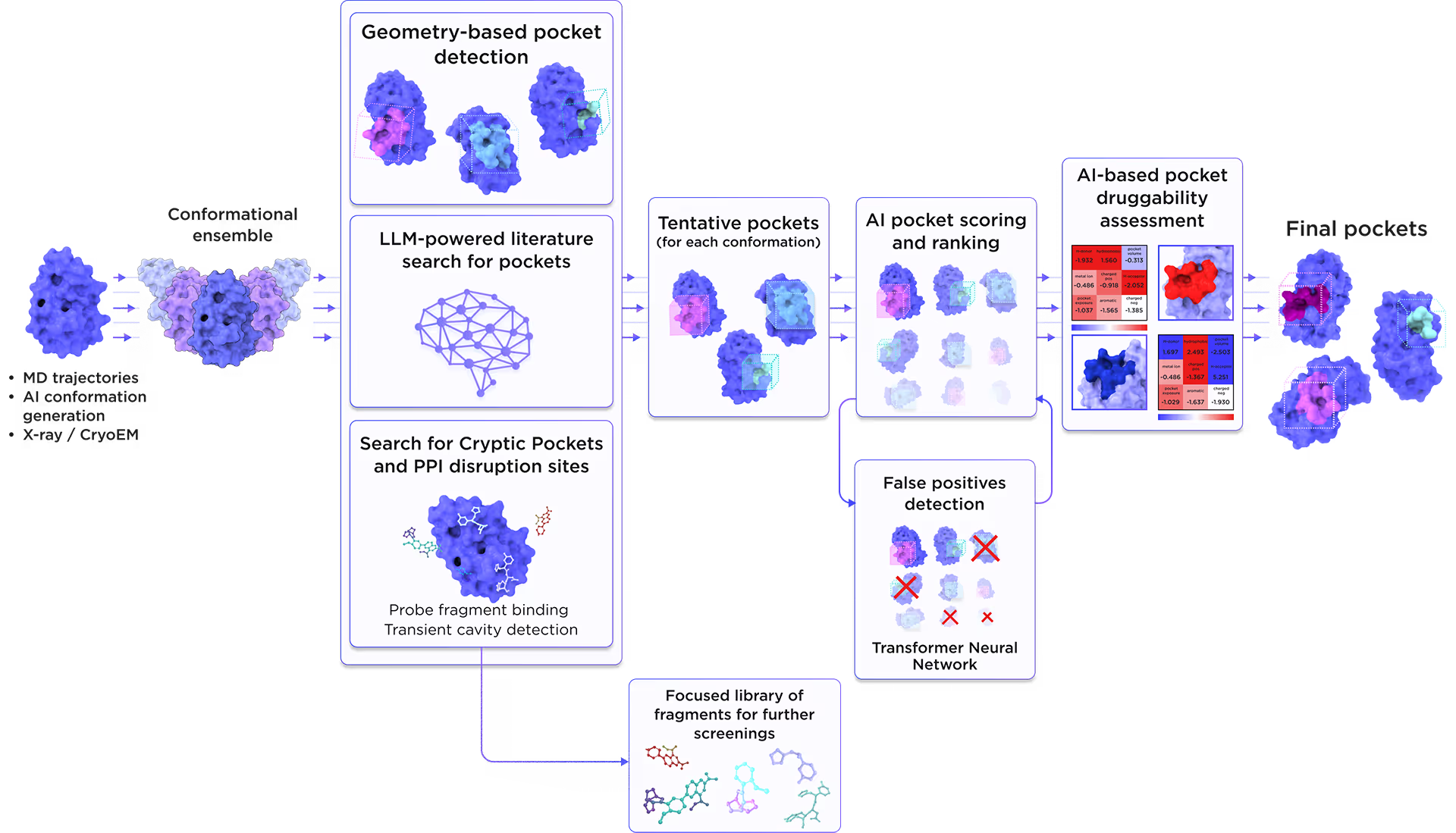
Fig. 3. AI-based binding pocket detection workflow
4. AI-Powered Virtual Screening
Our ecosystem is equipped to perform AI-driven virtual screening on Histone H3.3. With access to a vast chemical space and cutting-edge AI docking algorithms, we can rapidly and reliably predict the most promising, novel, diverse, potent, and safe small molecule ligands of Histone H3.3. This approach allows us to achieve an excellent hit rate and to identify compounds ready for advanced lead discovery and optimization.
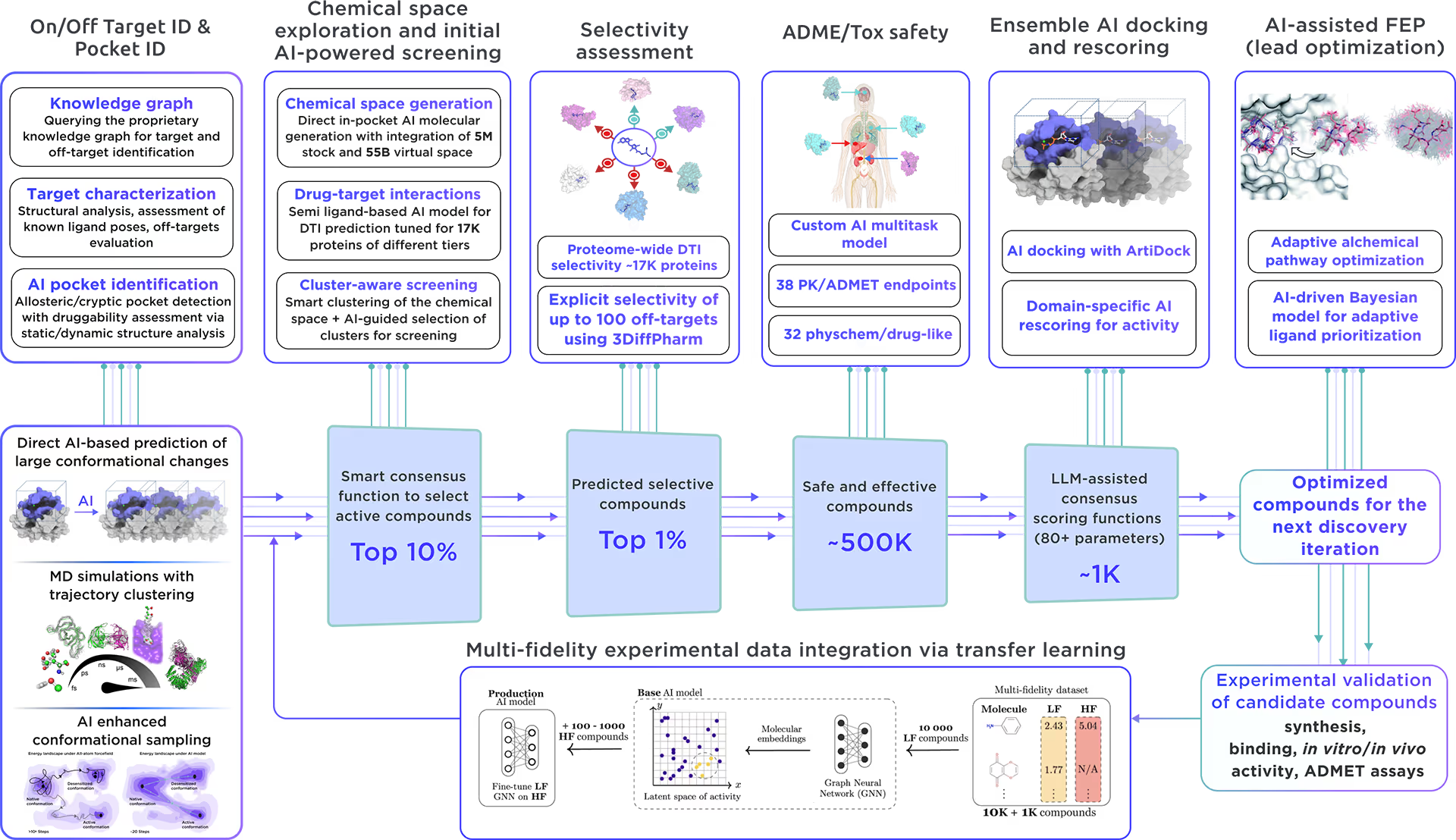
Fig. 4. The screening workflow of Receptor.AI
Receptor.AI, in partnership with Reaxense, developed a next-generation technology for on-demand focused library design to enable extensive target exploration.
The focused library for Histone H3.3 includes a list of the most effective modulators, each annotated with 38 ADME-Tox and 32 physicochemical and drug-likeness parameters. Furthermore, each compound is shown with its optimal docking poses, affinity scores, and activity scores, offering a detailed summary.
Histone H3.3
partner:
Reaxense
upacc:
P84243
UPID:
H33_HUMAN
Alternative names:
-
Alternative UPACC:
P84243; P06351; P33155; Q5VV55; Q5VV56; Q66I33; Q9V3W4
Background:
Histone H3.3, encoded by the gene with accession number P84243, is a variant histone that replaces conventional H3 in a wide range of nucleosomes in active genes. It is the predominant form of histone H3 in non-dividing cells and is incorporated into chromatin independently of DNA synthesis. This protein plays a crucial role in transcription regulation, DNA repair, DNA replication, and chromosomal stability through its involvement in chromatin structure and function.
Therapeutic significance:
Histone H3.3 is implicated in the pathogenesis of gliomas, including glioblastoma multiforme and diffuse intrinsic pontine glioma, through mutations that affect post-translational modifications. It is also associated with Bryant-Li-Bhoj neurodevelopmental syndrome 1 and 2, caused by variants in genes encoding this histone. Understanding the role of Histone H3.3 could open doors to potential therapeutic strategies for these conditions.