Balancing Affinity & Permeability
AI-Guided Multi-Objective Optimization of Peptides: Balancing Affinity & Permeability
Balancing Affinity & Permeability
Announcement



Full Text
The growing interest in peptide-based therapeutics stems largely from their natural strengths, including high target specificity compared with conventional small-molecule drugs. More recently, peptides have shown strong potential in targeting intracellular target proteins and protein-protein interactions that were considered undruggable by small molecules due to their shallow surface pockets.
However, peptides rarely achieve sufficient cellular membrane and gastrointestinal permeability, due to their large size, which directly impacts several structural properties, including high topological polar surface area (tPSA), many hydrogen-bond donors and acceptors, multiple ionizable groups, and high conformational flexibility. That is why designing peptides with high oral exposure that can reach intracellular targets remains one of the most important objectives in drug development.
Oral peptides in the Clinic
Despite these limitations, a small number of peptides have reached the clinic as oral drugs.
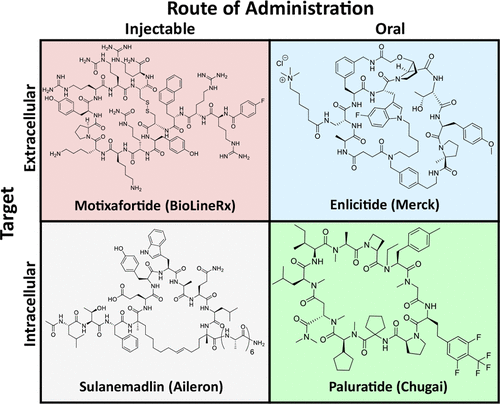
Enlicitide (Merck) is a prominent example of an extracellularly acting peptide, but it shows extremely low bioavailability and relies on co-formulation with a permeation enhancer to improve transport across the gastrointestinal epithelial layer. Another recent example is Paluratide (Chugai Pharmaceutical), which exhibits sufficiently high membrane permeability to reach an intracellular target. Other peptides, such as Sulanemadlin (Rein Therapeutics), that also act intracellularly, lack sufficient permeability to be classified as oral drugs and must be administered by injection.
Overall, most peptide discovery workflows optimize permeability through repeated cycles of design, synthesis, and experimental testing, creating a substantial laboratory demand as each new hypothesis must be validated empirically. Some of the key strategies for optimization include:
- Reducing exposed hydrogen bond donors and shielding heteroatoms
- Applying N-methylation, a validated approach for improving membrane permeability and oral bioavailability, particularly in cyclic peptides
- Promoting intramolecular hydrogen bonds to mask polar groups in nonpolar environments and facilitate membrane diffusion
- Constraining conformation through cyclization or stereochemical control to reduce entropic penalties
- Tuning conformational dynamics, often referred to as chameleonicity, so peptides remain sufficiently polar for solubility while adopting lower polarity conformations for membrane permeation
Peptide permeability is typically validated using experimental passive diffusion assays, such as PAMPA, and more biologically relevant cell based assays, such as Caco 2. The results from these assays are then interpreted to guide the next round of peptide optimization, including decisions about which structural features to preserve, modify, or deprioritize. The central limitation of this traditional workflow is the repeated back and forth between design and experiment, which requires intensive resources and becomes difficult to scale, especially when permeability depends on subtle conformational effects that are not readily apparent from assay data alone, with no other data source available.
In silico PAMPA
The simplest way to assess the passive membrane permeability of peptides is through PAMPA assay, which measures the rate of passive diffusion across an artificial lipid-infused membrane. Although PAMPA is relatively cheap, relying on it at every stage slows down the optimization procedure, since peptides have to be synthesized first and then tested.
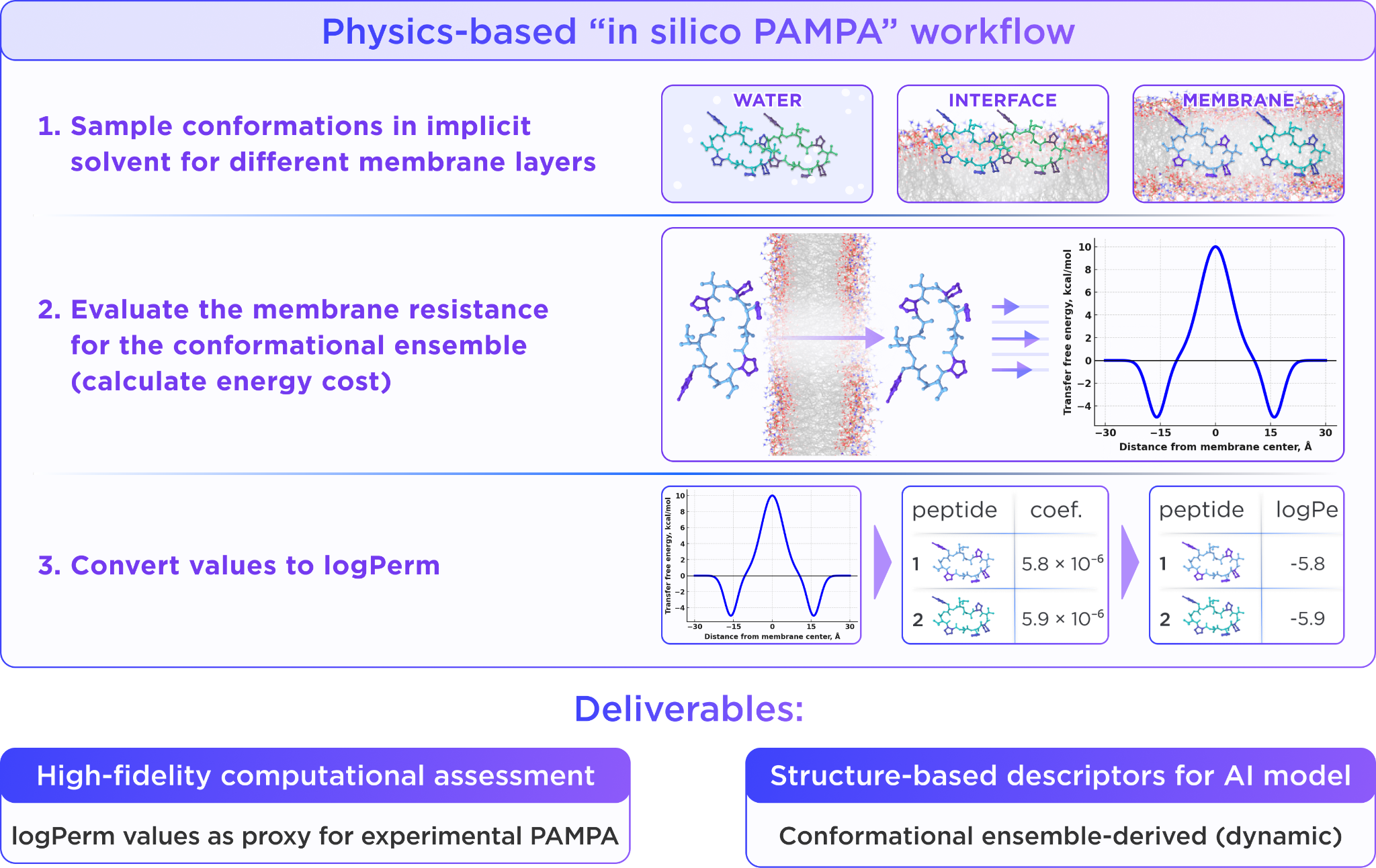
Therefore, at Receptor.AI, we built a physics-based computational workflow to generate a much faster permeability readout through three main steps. Initially, peptide conformations are sampled in aqueous and membrane environments, allowing the free energy cost of membrane transfer to be estimated across the full conformational ensemble. After evaluating membrane resistance, the resulting signal is converted into logPerm values, enabling high-fidelity permeability prediction and direct comparison with PAMPA measurements.
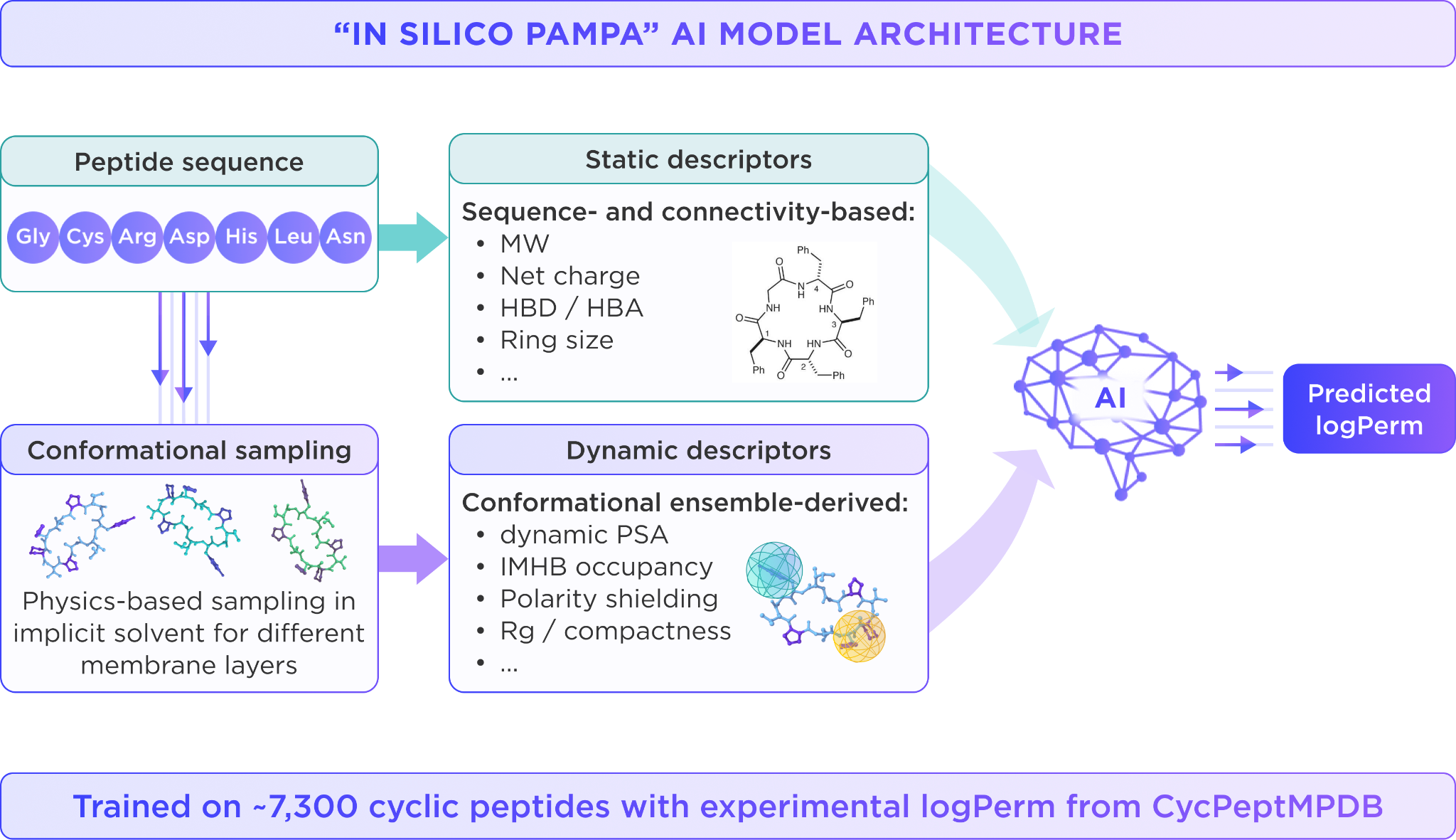
Additionally, we trained and validated an “in silico PAMPA” AI model with around 7300 cyclic peptides with experimental logPerm from CycPeptMPDB using random split. The model takes peptide sequences and static descriptors as inputs, then applies physics-based sampling to generate dynamic descriptors that determine permeability scores. Importantly, the model performs well not only on classical cyclic peptides, but also on lariat and lasso-like structures, suggesting that physics-based readouts improve generalization across diverse peptide topologies.
In practice, Receptor.AI’s “in silico PAMPA” model serves as a fast and scalable decision layer for permeability optimization within the broader AI platform.
Delta learning Engine
Moving from artificial membranes to cell-based in vitro assays provides a more relevant measure of cellular and GI permeability. Caco-2 assay is a prominent example and is widely used to model the permeability across the gastrointestinal tract, that can capture additional biological effects such as efflux and intracellular degradation.
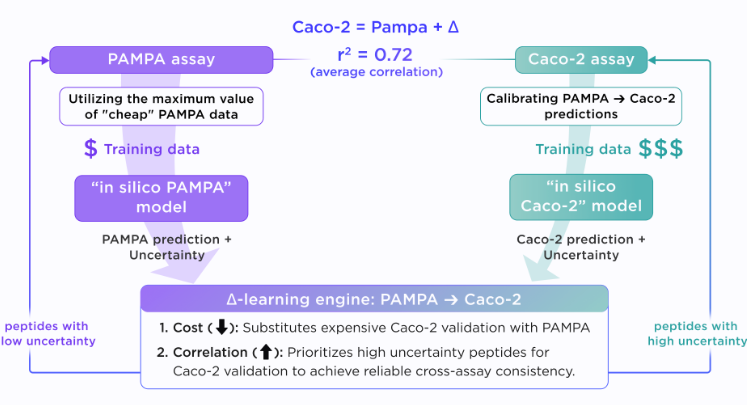
Thus, at Receptor.AI, alongside the “in silico PAMPA” model, we developed an in silico Caco-2 model trained on experimental Caco-2 assay data from CycPeptMPDB, enabling direct prediction of peptide permeability. Nevertheless, Caco-2 data are scarce and expensive to obtain, making it difficult to build scalable models from experimental measurements alone. To address this, we employ a delta-learning approach grounded in the observation that PAMPA and Caco 2 assay readouts are correlated, since both provide measures of passive permeability. The PAMPA ⇔ Caco-2 delta model is initially trained to learn the difference between PAMPA and observed Caco-2 measurements. Then, once a reliable in “silico PAMPA” prediction is available, the corresponding Caco-2 value can be estimated by adding the predicted delta. This allows PAMPA to be treated as a lower-fidelity but scalable anchor, and Caco-2 as a higher-fidelity assay that can be used selectively.
Effectively, the models are orchestrated by an AI decision engine that uses prediction uncertainty to determine which peptides should be prioritized for direct Caco-2 and ex-vivo validation. Consequently, expensive assays can be focused on the most informative peptides, accelerating model learning and reducing both cost and experimental burden.
Benchmarking
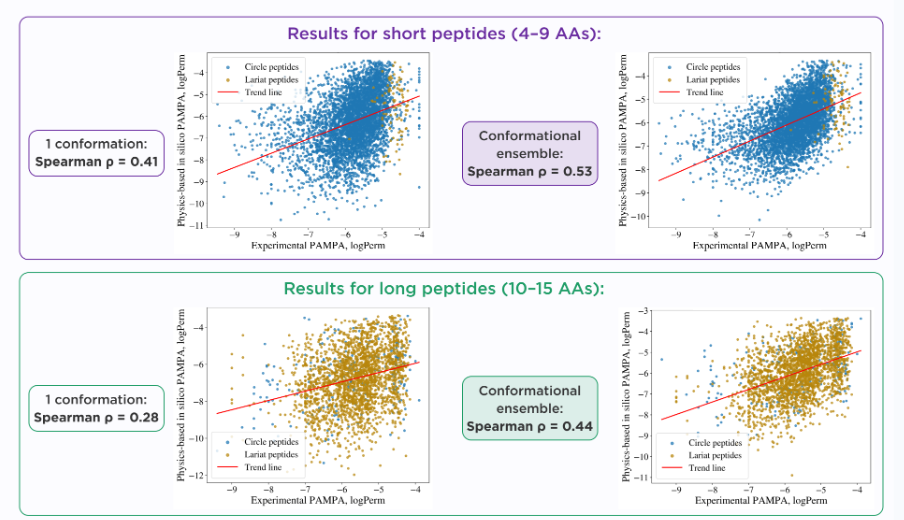
We benchmarked our physics-based workflow on two peptide subsets from CycPepMPDB: 4,503 short peptides (4-9 monomers) and 2,451 long peptides (10-15 monomers). As expected, the model showed lower correlation for the longer peptides, likely reflecting their greater structural complexity. However, in both subsets, correlation improved when conformational sampling was incorporated, as shown in the figure below.
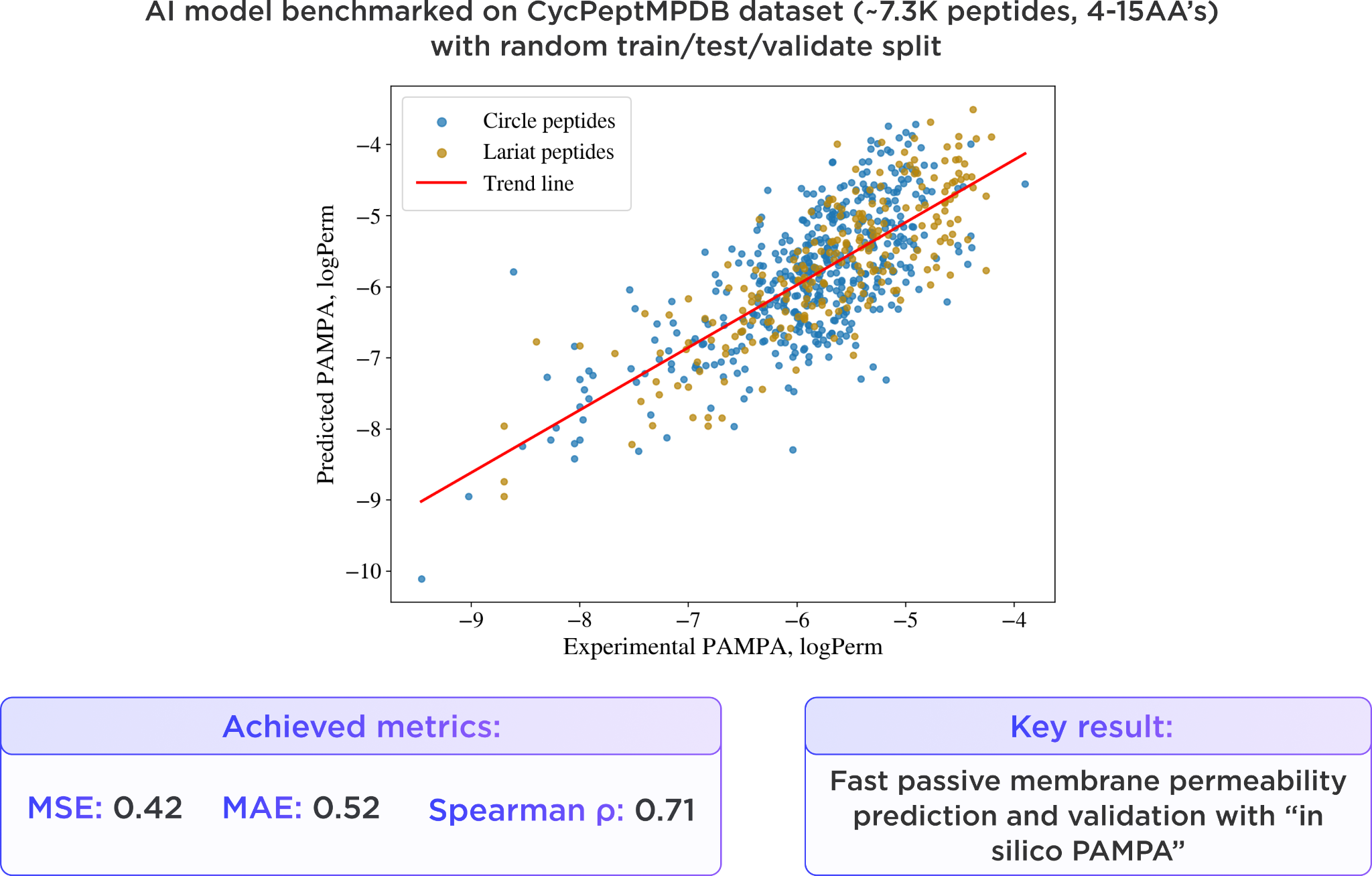
The same dataset was used to validate our “in silico PAMPA” AI model, which reproduced experimental PAMPA permeability trends with good overall accuracy and achieved a Spearman correlation of 0.71.
Similarly, our in silico Caco-2 model was trained on ~1300 cyclic peptides and demonstrated a high accuracy of 0.75.
In parallel, we trained our PAMPA-to-Caco-2 delta model on a matched dataset of approximately 230 cyclic peptides with both PAMPA and Caco-2 measurements. Even with this limited paired dataset, the model achieved a Spearman correlation of about 0.6 on the test set, demonstrating that a substantial part of the Caco-2 signal can be recovered by learning the PAMPA-to-Caco-2 delta.
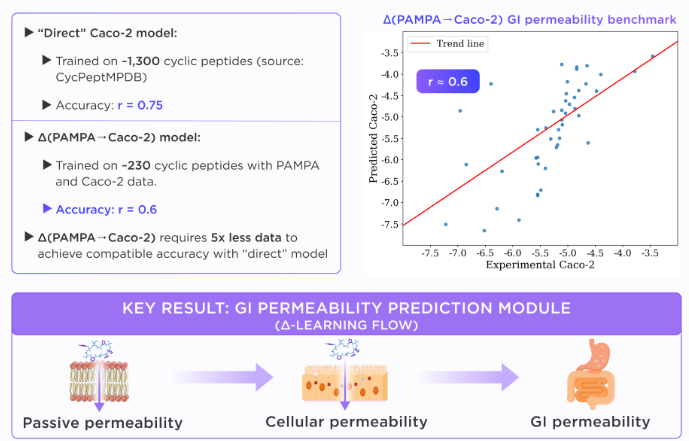
This workflow can be further expanded through our QuorumMap, which organizes computational and experimental methods into a closed learning system that supports multiparameter optimization of the properties required for oral exposure.
Case Study
In one of our case studies, we started from a 16-residue cyclic peptide targeting the intracellular PPI protein Vps29. The initial hit had moderate affinity, with a Kd of 780 nM, and poor membrane permeability, with a PAMPA logPerm of -8. We then compared two optimization strategies across the same search space of about 120 billion sequences, including non-canonical amino acids, while keeping the experimental setup identical.
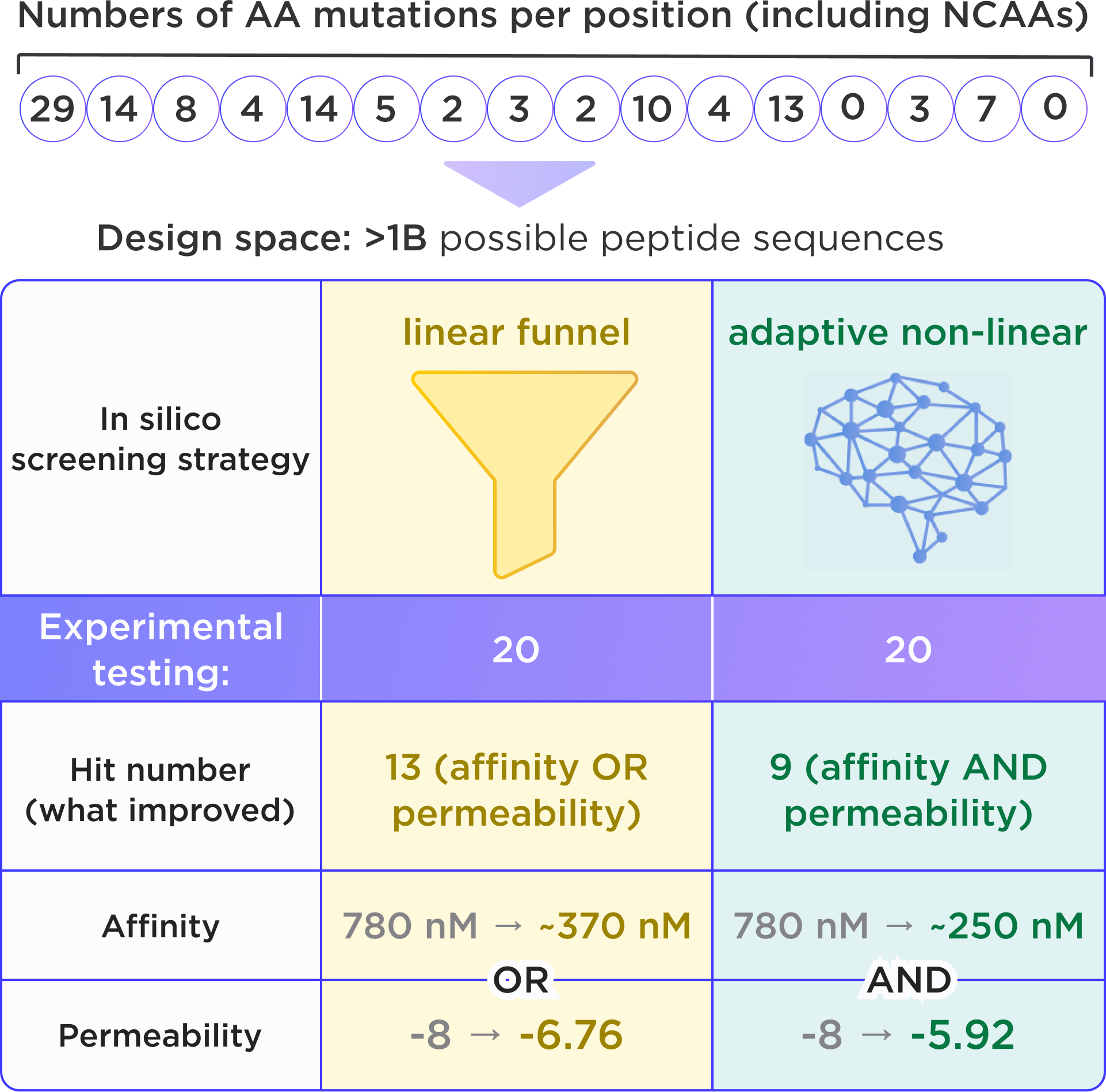
The difference in outcome was clear. Linear screening produced 13 candidates, but improvements were limited to a single property: the best peptide achieved only a 2-fold gain in affinity, while the most permeable compound reached logPerm = -6.8 at the cost of weaker binding. In contrast, AI-orchestrated non-linear screening produced 10 candidates with simultaneous gains in both affinity and permeability, with the top peptide delivering a 3-fold improvement in affinity and improving PAMPA permeability to about -6.
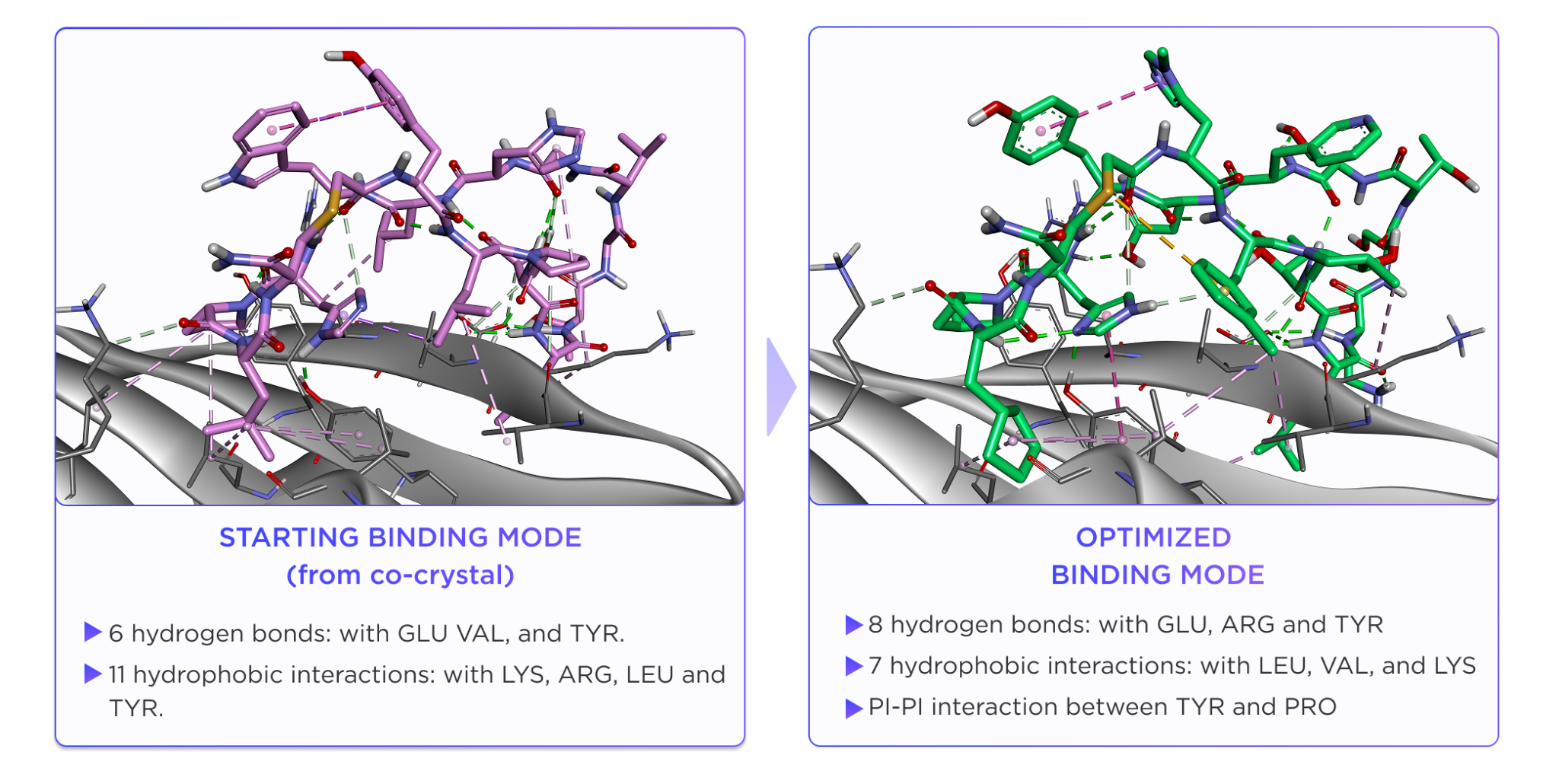
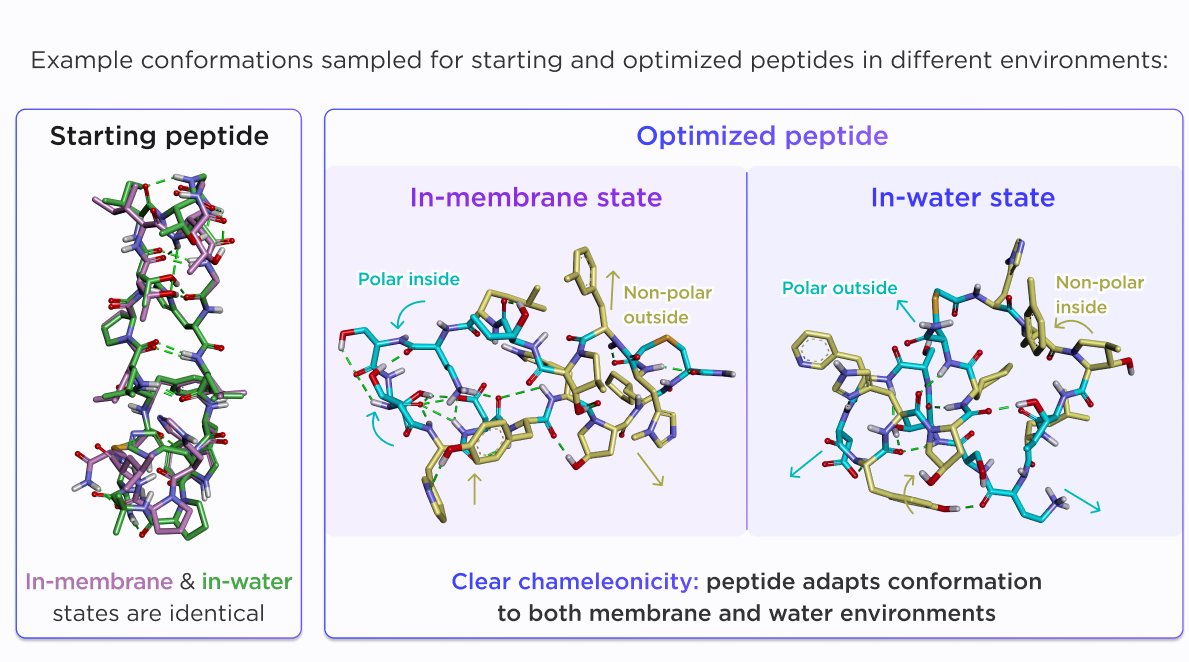
As a result, the optimized peptide showed greater conformational adaptability across environments. In membrane-like conditions, hydrophobic residues remained exposed and polar groups were buried through an internal hydrogen-bond network, while in water the peptide reorganized to reduce hydrophobic surface exposure. By contrast, the starting peptide remained locked in essentially the same rigid beta-sheet-like conformation in both environments.
Summary
Peptide permeability remains one of the main bottlenecks in developing orally available peptide therapeutics. To address this, at Receptor.AI, we built a physics-based “in silico PAMPA” workflow that simulates membrane-crossing energetics and translates them into a permeability readout, directly comparable to experimental PAMPA. This framework is extended through a PAMPA↔Caco-2 delta-learning engine, enabling scalable prediction of higher-fidelity cellular and GI permeability and reducing the amount of required Caco-2 or ex vivo experiments. To support oral bioavailability optimization, this framework is orchestrated with the QuorumMap decision engine, enabling permeability to be optimized alongside other key drug properties within a unified decision-making workflow.

