
Predicting Peptide Permeability at Scale
Announcement



Full Text
Poor membrane permeability remains one of the main reasons peptide programs stall before reaching intracellular targets. While medicinal chemistry has made substantial progress in improving potency and stability, permeability is still too often assessed late, after synthesis, when options are already constrained.
At Receptor.AI, we approach peptide permeability as a design-time property, not a post hoc filter. Our focus is on passive membrane permeability: the component most amenable to mechanistic modeling and early optimization, and one that imposes a strong practical constraint on what downstream formulation or delivery strategies can realistically compensate for.
This post outlines how our physics-based permeability workflow operates, how it is benchmarked, and how it integrates directly into peptide optimization.
Why peptide permeability resists simple rules
Unlike small molecules, peptides exist as conformational ensembles, continuously reorganizing their polar surface, hydrogen-bonding patterns, and compactness across aqueous and membrane environments. As a result, familiar descriptors such as logP, polar surface area, or molecular weight fail to capture key determinants of permeability.
This challenge is amplified in macrocyclic and constrained peptides, where chameleonic behavior allows polar groups to be transiently shielded in membranes while remaining exposed in water. Capturing this behavior requires models that explicitly account for structure, dynamics, and membrane interaction, rather than statistics over fixed descriptors.
A physics-based view of passive diffusion
Receptor.AI’s permeability workflow is built on the premise that passive membrane permeability is governed by the free-energy cost of crossing a lipid bilayer, taking into account the conformational states a peptide can realistically adopt.
In practice, this involves generating a peptide conformational ensemble, simulating membrane insertion and translocation using atomistic models and continuum electrostatics, computing free-energy profiles across the membrane, and converting these profiles into intrinsic permeability coefficients that reflect passive diffusion behavior.
At the free-energy level, permeability estimation is fully physics-derived; no experimental permeability values are introduced at this stage.

From free energy to usable design objectives
Intrinsic permeability coefficients are not directly actionable for design. To integrate physics-derived estimates into optimization workflows, Receptor.AI rescales these values onto a Caco-2–like scale using correlations with experimental data, providing a calibrated interface between mechanistic simulation output and experimental intuition without overwriting the underlying physics.
Experimental PAMPA or Caco-2 measurements enter the workflow only after selected designs are synthesized and tested. These data are treated as separate signals within the optimization environment, updating priorities and guiding subsequent search rather than being blended into every prediction step.
Benchmarking at scale
The workflow was benchmarked on ~8,000 cyclic peptides annotated with experimental PAMPA and Caco-2 permeability data. Predicted permeability showed strong rank correlation with PAMPA (Spearman ρ = 0.83) and more modest correlation with Caco-2 (ρ = 0.51). For context, PAMPA and Caco-2 themselves correlate at ~ρ = 0.57 for peptides.
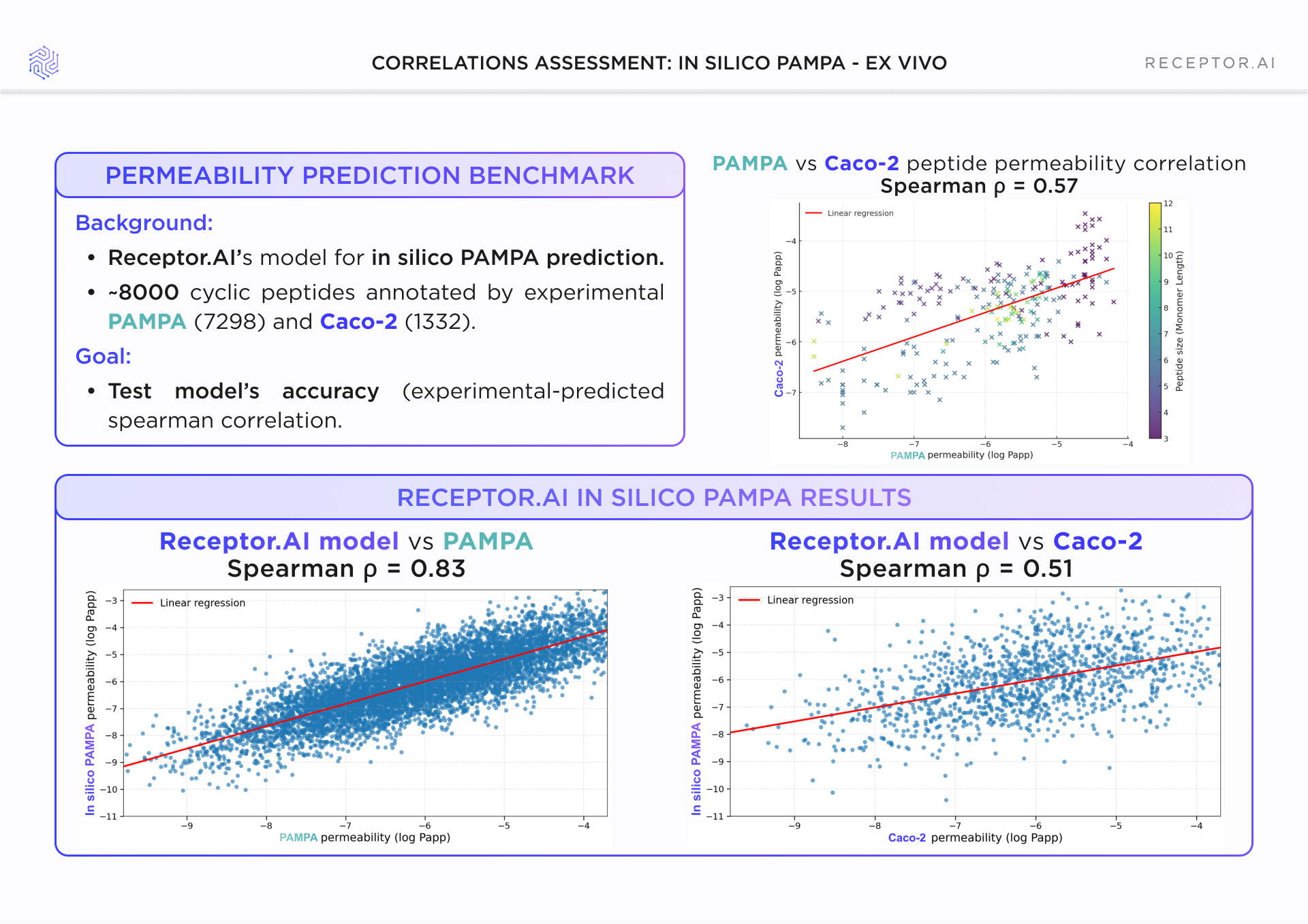
These results place the physics-based workflow within the intrinsic variability of commonly used permeability assays. The output is best interpreted as a prioritization and filtering signal, rather than a precise numerical predictor.
Permeability as part of multi-parameter optimization
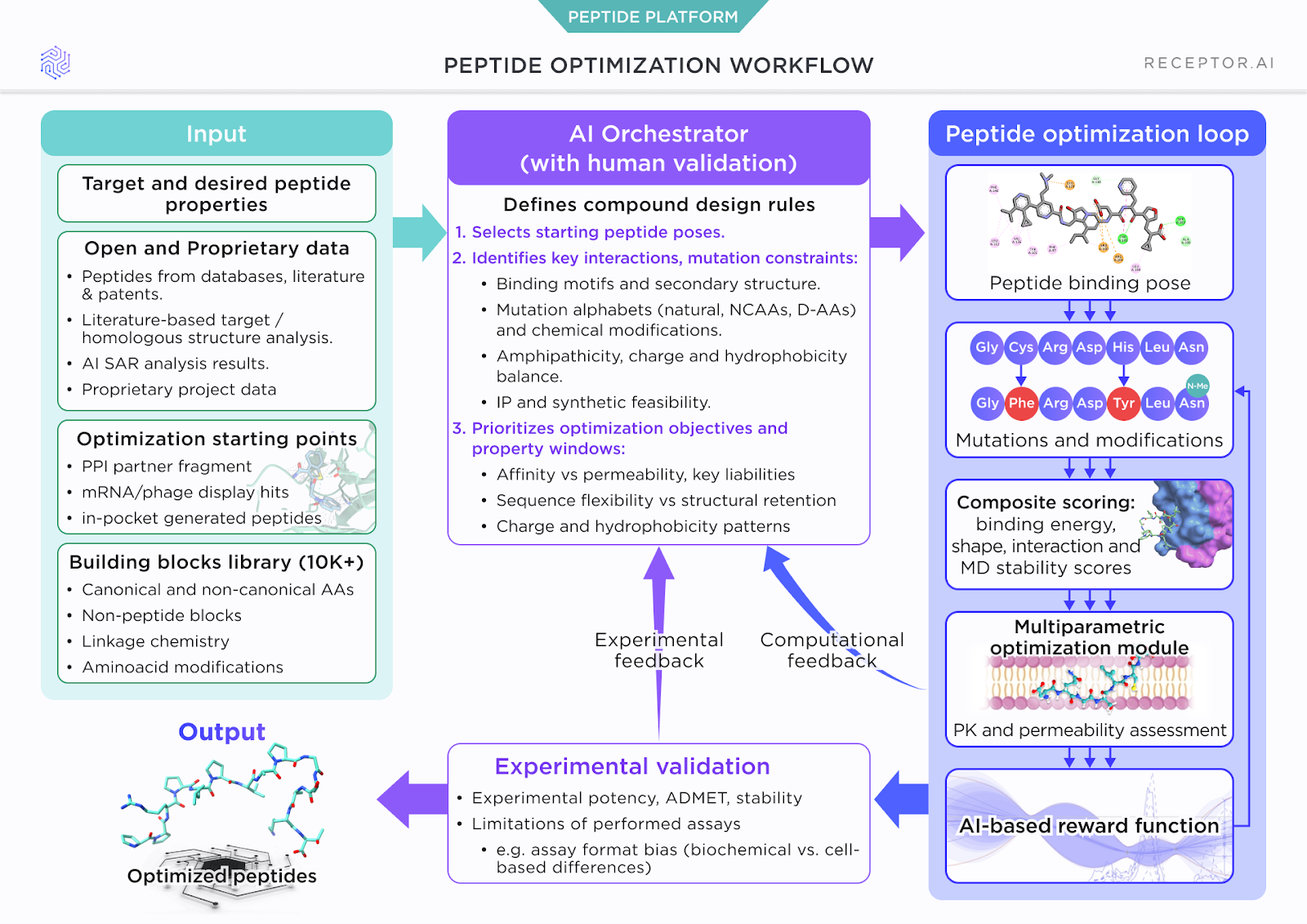
Permeability prediction delivers its full value only when embedded directly into peptide design. Within Receptor.AI’s optimization framework, permeability is evaluated alongside binding affinity, structural stability, and sequence feasibility.
An AI orchestrator defines allowed mutations, including D- and non-canonical residues, enforces sequence–structure constraints, and balances competing objectives through a reward function. This enables a 10–100× reduction in candidate space, far beyond what manual SAR or intuition can reasonably explore, while experimental feedback on activity and permeability continuously refines the design loop.
From passive permeability to exposure
Passive permeability is not equivalent to oral bioavailability. Transporters, efflux, metabolism, and formulation all play critical roles downstream. However, across peptide datasets, passive permeability correlates with intestinal permeability, and intestinal permeability correlates more weakly with oral exposure. Peptides with vanishing passive permeability rarely succeed through downstream intervention alone.
For highly potent peptides, even modest improvements in permeability, shifting absorption from ~1–2% to ~3–10%, can be sufficient for efficacy. This makes early, physics-based permeability optimization a practical leverage point, even though it is not a guarantee of oral success.
By grounding permeability estimation in physical principles and integrating it directly into peptide optimization, Receptor.AI shifts permeability from an empirical afterthought to a quantitative design constraint. The goal is not to replace PAMPA, Caco-2, or in vivo studies, but to prioritize synthesis by focusing experimental effort on peptides with a realistic chance of crossing membranes efficiently.
Learn more about PEPTOR™, Receptor.AI’s platform for peptide design

